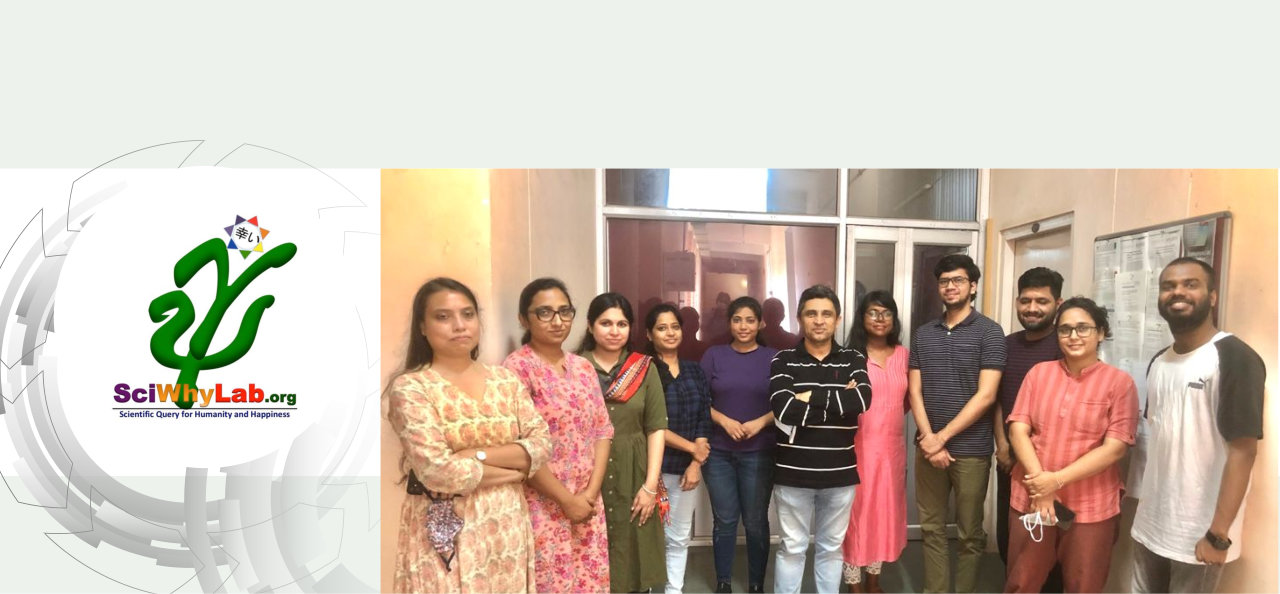
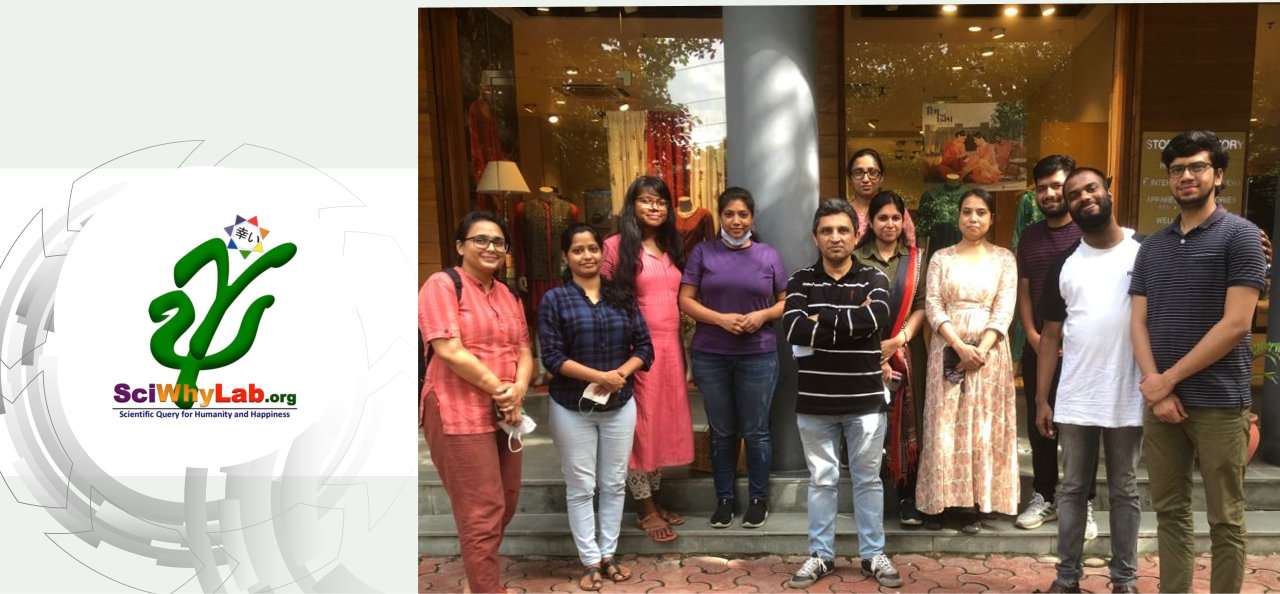
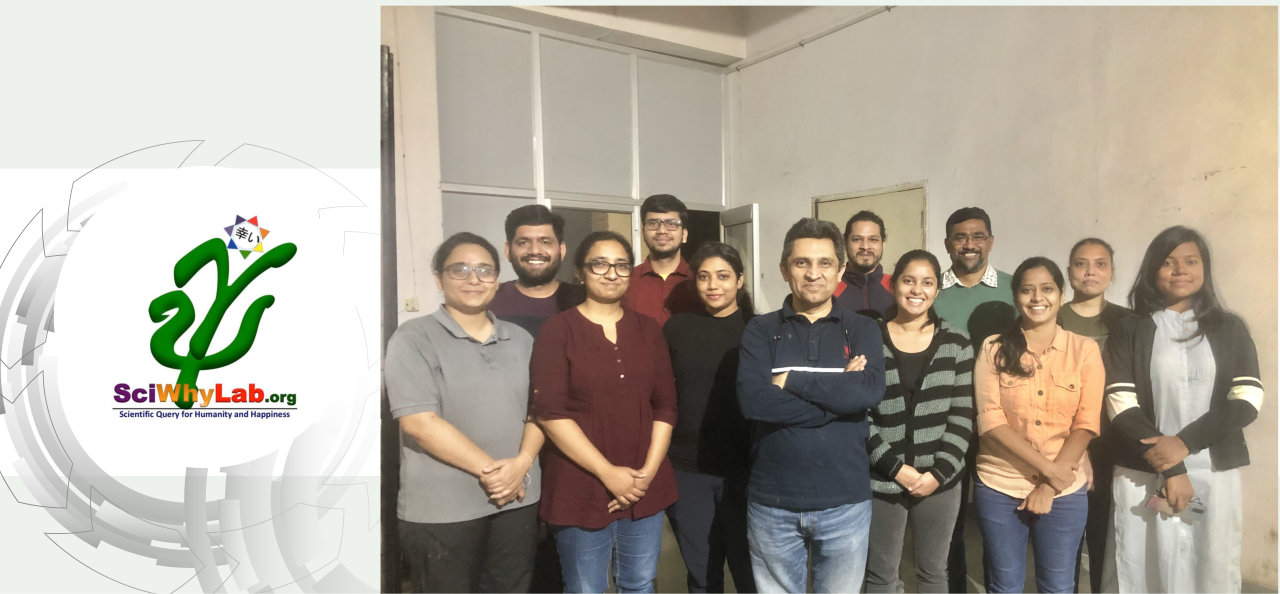
About Us
We are located at School of Computational & Integrative Sciences, Jawaharlal Nehru University (JNU), New Delhi. JNU is a prestigious Indian university, recently ranked at no. 3 in the country as per the Government instituted committee for this purpose. School of Computational and Integrative Sciences is a relatively young institution with JNU with its thrust research areas being Bioinformatics, Complex Systems and Big data analytics.
SciWhyLab stands for scientific query for human health and happiness. The word also draws on the pun with the Japanese word 幸い(sai-wai) which implies deriving "happiness" from our work.
We are interested in developing data-driven algorithms and applications for biological data with an integrative perspective. We aim to leverage on our past and ongoing works in the field of sequence and structural analysis of protein-DNA and other biomolecular interactions, particularly their modelling through machine learning algorithms. Our technical research interests are in machine learning, big data analytics and novel architectures and training methods for deep learning through neural networks.
Our current projects involve a broad range of biological data science problems, such as DNA shape based transcriptional recognition, single cell transcriptomics, transcriptome dimensionality reduction, medical diagnosis from clinical data, genome-phenome associations and we address most of these problems with a data-driven approach at genomic scale. We are collaborating with academia and industry in the field of head & neck cancer, Myobacterium Tuberculosis therapeutic target discovery, pre-natal non-invasive diagnostics and Biomarker discovery.
We belive in creativity, rigour and excellence and constantly strive to improve ourselves. Suggestions and comments may be directly sent to the group leader.
 Prof. Shandar Ahmad
Prof. Shandar Ahmad
As a Principal Investigator of this group: Science, poetry and pretentious thinking are all my passions, of which I chose the first as a profession. I do science for the artist's vanity and a decent career it can paradoxically combine to offer. It is delusional enough to make me lose sight of the trivial world, and useful enough so I can pay my bills and lead a happy life. This website is a public and semi-personal portal, to advertise myself and my group (under development) and put together a summary of our achievements in the course of our random walk across the beautiful and unusual forays offered by our deliberate and chance encounters with knowledge.









